Marshall Lab Research
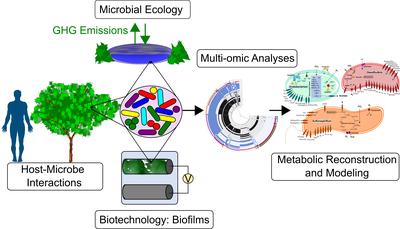
The Marshall Lab at Marquette University is an interdisciplinary team of microbiologists, biochemists, bioinformaticians, and engineers. We are interested in many areas of microbial ecology and evolution, including the pathogen adaptation in host environments, evolution of antibiotic resistance, bioelectrochemistry, bioremediation, and genomics.